|
Chang, Y. W., et al., Oncogene 33.43 (2014): 5065-5077.
Mai, Ru-Tsun, et al., American Journal of Cancer Research 12.12 (2022): 5462.
|
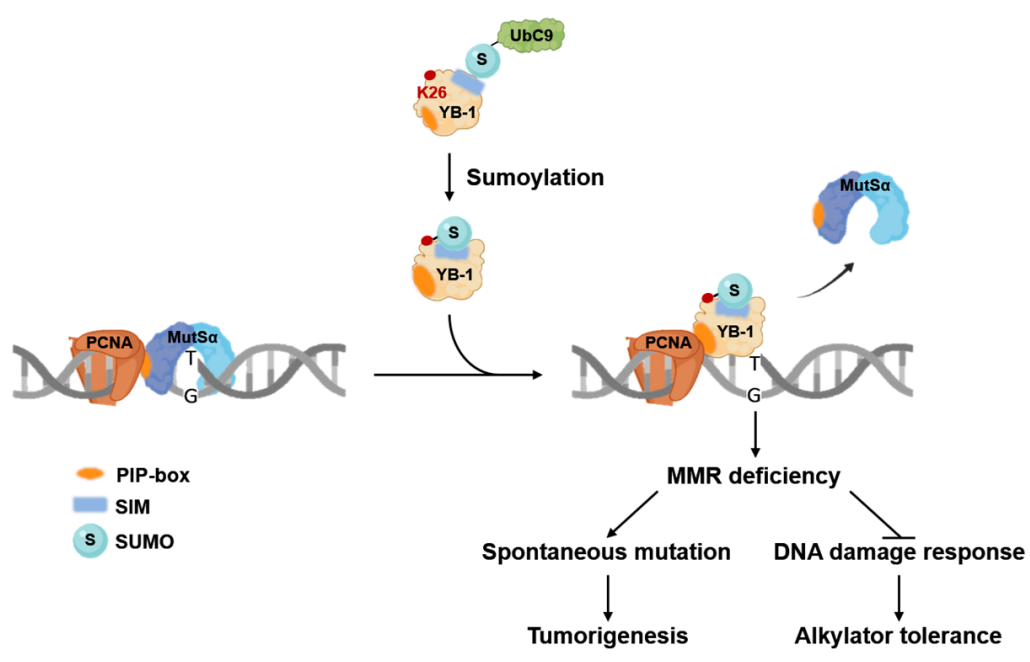
Sumoylation Participates in the Regulation of YB-1-mediated Mismatch Repair Deficiency and Alkylator Tolerance
Numerous reports indicate that enhanced expression of Y-box binding protein-1 (YB-1) in tumor cells is strongly associated with tumorigenesis, aggressiveness, drug resistance, as well as poor prognosis in several types of cancers, and YB-1 is considered to be an oncogene. The molecular mechanism contributing to the regulation of the biological activities of YB-1 remains obscure. Sumoylation, a post-translational modification involving the covalent conjugation of small ubiquitin-like modifier (SUMO) proteins to a target protein, plays key roles in the modulation of protein functions. In this study, our results revealed that YB-1 is sumoylated and that Lys26 is a critical residue for YB-1 sumoylation....
YB-1 disrupts Mismatch Repair Complex Formation, Interferes with MutSα Recruitment on Mismatch and Inhibits Mismatch Repair through Interacting with PCNA
Y-box binding protein-1 (YB-1) is highly expressed in tumors and it participates in various cellular processes. Previous studies indicated that YB-1 binds to mispaired DNA and interacts with several mismatch repair (MMR)-related factors. However, its role in the MMR system remains undefined. Here, we found that YB-1 represses mutS homolog 6 (MSH6)-containing MMR complex formation and reduces MutSα mismatch binding activity by disrupting interactions among MMR-related factors. In an effort to elucidate how YB-1 exerts this inhibitory effect, we have identified two functional proliferating cell nuclear antigen (PCNA)-interacting protein (PIP)-boxes that mediate YB-1/PCNA interaction and locate within the C-terminal region of YB-1...
|
|
Su, Yi-Yuan, et al., American Journal of Cancer Research 13.5 (2023): 1744.
|
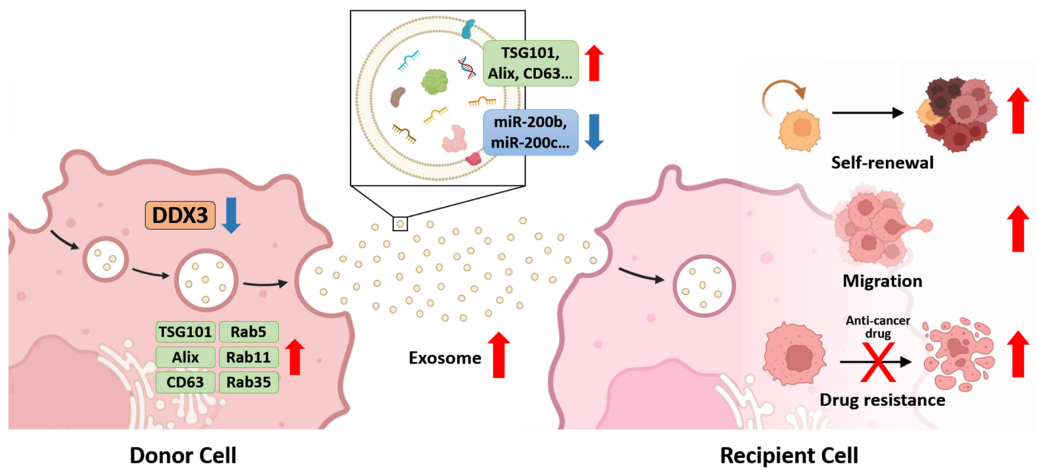
DDX3 Suppresses Hepatocellular Carcinoma Progression Through Modulating the Secretion and Composition of Exosome
Due to the lack of predictive biomarkers and the lack of conspicuous symptoms at the early stage, hepatocellular carcinoma (HCC) remains difficult to diagnose and treat effectively. During cancer development, exosomes secreted from tumor cells carry functional molecules to surrounding recipient cells, thereby participating in the regulation of cancer progression. DDX3, a DEAD-box RNA helicase, performs many important functions in several cellular processes and is therefore implicated as a tumor suppressor in HCC. However, whether DDX3 affects the secretion and cargo sorting of HCC exosomes remains obscure. In this study, our results revealed that reduced DDX3 expression in HCC cells promoted the release of exosomes and enhanced the expression of several exosome biogenesis-associated proteins....
|
|
Li, Hao-Kang, et al., Scientific reports 6.1 (2016): 28637.
|
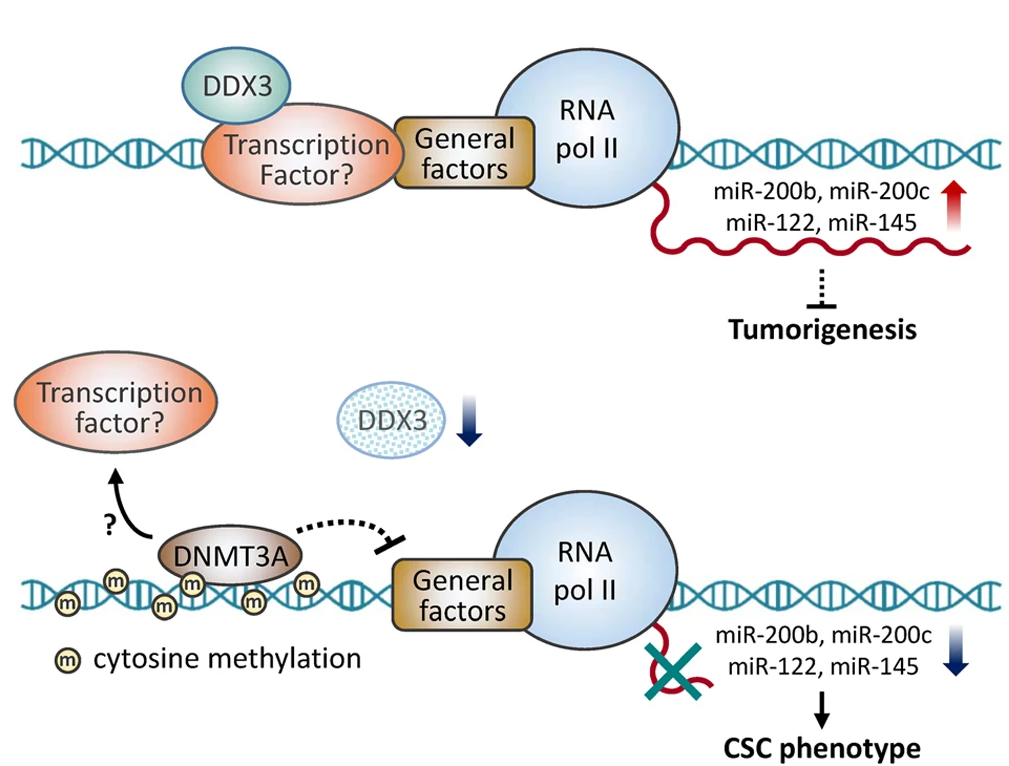
DDX3 Represses Stemness by Epigenetically Modulating Tumor-suppressive miRNAs in Hepatocellular Carcinoma
Studies indicate that the presence of cancer stem cells (CSCs) is responsible for poor prognosis of hepatocellular carcinoma (HCC) patients. In this study, the functional role of DDX3 in regulation of hepatic CSCs was investigated. Our results demonstrated that reduced DDX3 expression was not only inversely associated with tumor grade, but also predicted poor prognosis of HCC patients. Knockdown of DDX3 in HCC cell line HepG2 induced stemness gene signature followed by occurrence of self-renewal, chemoreisistance, EMT, migration as well as CSC expansion, and most importantly, DDX3 knockdown promotes tumorigenesis. Moreover, we found positive correlations between DDX3 level and expressions of tumor-suppressive miR-200b, miR-200c, miR-122 and miR-145, but not miR-10b and miR-519a, implying their involvement in DDX3 knockdown-induced CSC phenotypes...
|
Publication List
1. Su, Y.-Y., Chao, C.-H., Hsu, H.-Y., Li, H.-K., Wang, Y.-L., Lee, Y.-H. W., and Mai, R.-T. (2023, Apr). DDX3 suppresse hepatocellular
carcinoma progression through modulating the secretion and composition of exosome. Am. J. Cancer Res., 13(5):1744-1765.
2. Mai, R.-T., Chao, C.-H., Chang, Y.-W., Kao, Y.-C., Cheng, Y, Hsu, H.-Y., Su, Y.-Y., Wang, C.-Y., and Lai, B.-Y. (2022, Dec).
Sumoylation participates in the regulation of YB-1-mediated mismatch repair deficiency and alkylator tolerance. Am. J.
Cancer Res., 12(12):5462-5483.
3. Wang, C.-Y., Wang, C.-H., Mai, R.-T., Chen T.-W., Li C.-W., and Chao, C.-H. (2022, Jul). Mutant p53-microRNA-200c-ZEB2-axis
induced CPT1C elevation contributes to metabolic reprogramming and tumor progression in basal-like breast
cancers.Front. Oncol., 12:940402.
4. Chao, C.-H., Wang, C.-Y., Wang, C.-H., Chen T.-W., Hsu, H.-Y., Huang, H.-W., Li, C.-W., and Mai, R.-T. (2021, Nov). Mutant p53
attenuates oxidative phosphorylation and facilitates cancer stemness through downregulating miR-200c-PCK2 axis in
basal like breast cancer. Mol. Cancer Res., 19(11):1900-1916.
5. Hsu P.-H., Chen W.-H., Juan-Lu C., Hsieh S.-C., Lin S.-C., Mai R.-T., and Chen S.-Y. (2021, Sep). Hesperidin and chlorogenic acid
synergistically inhibit the growth of breast cancer cells via estrogen receptor/mitochondrial pathway. Life (Basel),
11(9):950.
6. Li, H.-K., Mai, R.-T., Huang, H.-D., Chou, C.-H., Chang, Y.-A., Chang, Y.-W., You, L.-R., Chen, C.-M., and Lee, Y.-H. W. (2016, Jun).
DDX3 represses stemness by epigenetically modulating tumor-suppressive miRNAs in hepatocellular carcinoma. Sci. Rep.,
6:28637.
7. Chang, Y.-W., Mai, R.-T., Fang, W.-H., Lin, C.-C., Chiu, C.-C., and Lee, Y.-H. W. (2014, Oct). YB-1 disrupts mismatch repair
complex formation, interferes with MutSα recruitment on mismatch and inhibits mismatch repair through interacting
with PCNA. Oncogene, 33(43):5065-5077.
8. Huang, W.-H., Mai, R.-T., and Lee, Y.-H. W. (2008). Transcription factor YY1 and its associated acetyltransferases CBP and
p300 interact with hepatitis delta antigens and modulate hepatitis delta virus RNA replication. J. Virol. 82, 7313-7324.
9. Liao, W.-R., Hsieh, R.-H., Hsu, K.-W., Wu, M.-Z., Tseng, M.-J., Mai, R.-T., Lee, Y.-H. W., and Yeh, T.-S. (2007). The CBF1 -
independent Notch1 signal pathway activates human c-myc expression partially via transcription factor
YY1. Carcinogenesis28, 1867-1876.
10. Mai, R.-T., Yeh, T.-S., Kao, C.-F., Sun, S.-K., Huang, H.-H., and Lee, Y.-H. W. (2006). Hepatitis C virus core protein recruits
nucleolar phosphoprotein B23 and co-activator p300 to relieve the repression effect of transcriptional factor YY1 on B23
gene expression. Oncogene 25, 448-462.
11. You, L.-R., Chen, C.-M., Yeh, T.-S., Tsai, T.-Y., Mai, R.-T., Lin, C.-H., and Lee, Y.-H. W. (1999). Hepatitis C virus core protein
interacts with cellular putative RNA helicase. J. Virol. 73, 2841-2853.
carcinoma progression through modulating the secretion and composition of exosome. Am. J. Cancer Res., 13(5):1744-1765.
2. Mai, R.-T., Chao, C.-H., Chang, Y.-W., Kao, Y.-C., Cheng, Y, Hsu, H.-Y., Su, Y.-Y., Wang, C.-Y., and Lai, B.-Y. (2022, Dec).
Sumoylation participates in the regulation of YB-1-mediated mismatch repair deficiency and alkylator tolerance. Am. J.
Cancer Res., 12(12):5462-5483.
3. Wang, C.-Y., Wang, C.-H., Mai, R.-T., Chen T.-W., Li C.-W., and Chao, C.-H. (2022, Jul). Mutant p53-microRNA-200c-ZEB2-axis
induced CPT1C elevation contributes to metabolic reprogramming and tumor progression in basal-like breast
cancers.Front. Oncol., 12:940402.
4. Chao, C.-H., Wang, C.-Y., Wang, C.-H., Chen T.-W., Hsu, H.-Y., Huang, H.-W., Li, C.-W., and Mai, R.-T. (2021, Nov). Mutant p53
attenuates oxidative phosphorylation and facilitates cancer stemness through downregulating miR-200c-PCK2 axis in
basal like breast cancer. Mol. Cancer Res., 19(11):1900-1916.
5. Hsu P.-H., Chen W.-H., Juan-Lu C., Hsieh S.-C., Lin S.-C., Mai R.-T., and Chen S.-Y. (2021, Sep). Hesperidin and chlorogenic acid
synergistically inhibit the growth of breast cancer cells via estrogen receptor/mitochondrial pathway. Life (Basel),
11(9):950.
6. Li, H.-K., Mai, R.-T., Huang, H.-D., Chou, C.-H., Chang, Y.-A., Chang, Y.-W., You, L.-R., Chen, C.-M., and Lee, Y.-H. W. (2016, Jun).
DDX3 represses stemness by epigenetically modulating tumor-suppressive miRNAs in hepatocellular carcinoma. Sci. Rep.,
6:28637.
7. Chang, Y.-W., Mai, R.-T., Fang, W.-H., Lin, C.-C., Chiu, C.-C., and Lee, Y.-H. W. (2014, Oct). YB-1 disrupts mismatch repair
complex formation, interferes with MutSα recruitment on mismatch and inhibits mismatch repair through interacting
with PCNA. Oncogene, 33(43):5065-5077.
8. Huang, W.-H., Mai, R.-T., and Lee, Y.-H. W. (2008). Transcription factor YY1 and its associated acetyltransferases CBP and
p300 interact with hepatitis delta antigens and modulate hepatitis delta virus RNA replication. J. Virol. 82, 7313-7324.
9. Liao, W.-R., Hsieh, R.-H., Hsu, K.-W., Wu, M.-Z., Tseng, M.-J., Mai, R.-T., Lee, Y.-H. W., and Yeh, T.-S. (2007). The CBF1 -
independent Notch1 signal pathway activates human c-myc expression partially via transcription factor
YY1. Carcinogenesis28, 1867-1876.
10. Mai, R.-T., Yeh, T.-S., Kao, C.-F., Sun, S.-K., Huang, H.-H., and Lee, Y.-H. W. (2006). Hepatitis C virus core protein recruits
nucleolar phosphoprotein B23 and co-activator p300 to relieve the repression effect of transcriptional factor YY1 on B23
gene expression. Oncogene 25, 448-462.
11. You, L.-R., Chen, C.-M., Yeh, T.-S., Tsai, T.-Y., Mai, R.-T., Lin, C.-H., and Lee, Y.-H. W. (1999). Hepatitis C virus core protein
interacts with cellular putative RNA helicase. J. Virol. 73, 2841-2853.